
iris-r-gateway-template  Issue Detected
Issue Detected


 4
4 0
0
What's new in this version
Initial Release
iris-r-gateway-template
This repository is a template for using the new R gateway for IRIS 2021.1.
The architecture of this template is as follows:
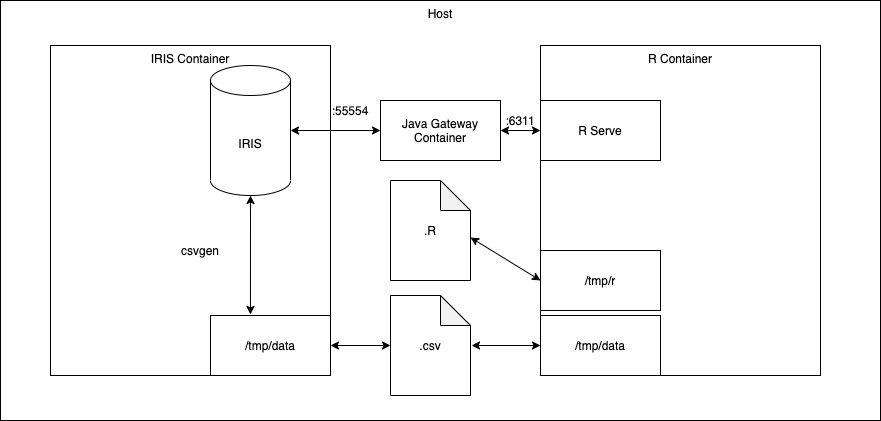
It is composed of three containers, with one for:
- IRIS
- R and RServe
- A Java Gateway
Iris and the R server will discuss through the java gateway.
These first two containers are bound to the data/ local folder, and so have access to the same data:
- abalone.csv
This file is loaded as a table in IRIS :
- Test.Data
The R container also has access to the r/ folder, and in it is the following script:
- test.R
getMode <- function(x) { l <- unique(x) l[which.max(tabulate(match(x, l)))] }getLength <- function(x) { length(x) }
createHistJPG <- function(data, dir, label) { dir2 = paste(dir, "/hist.jpeg", sep="") jpeg(file = dir2) hist(data, xlab=label) dev.off() }
createHistPNG <- function(data, dir, label) { dir2 = paste(dir, "/hist.png", sep="") png(file = dir2) hist(data, xlab=label) dev.off() }
createHistPDF <- function(data, dir, label) { dir2 = paste(dir, "/hist.pdf", sep="") pdf(file = dir2) hist(data, xlab=label) dev.off() }
Run this template
git clone https://github.com/grongierisc/iris-r-gateway-template
then :
docker compose up
Play with this template
Run the following ObjectScript commands
Run random R expression
do ##class(Demo.RGateway).RunEval()
output :
"R version 3.4.4 (2018-03-15)"
code :
ClassMethod RunEval() As %Status { // Get a gateway instance set gateway = ##class(%Net.Remote.Gateway).%New() set tSC = gateway.%Connect(..#JGWHOST, ..#JGWPORT) if $$$ISERR(tSC) quit// Bind the gateway to the Rserve set c = ##class(%Net.Remote.Object).%ClassMethod(.gateway,"com.intersystems.rgateway.Helper","createRConnection", ..#RHOST, ..#RPORT) // Run a random R command zw c.eval("R.version$version.string").asString() do:c'="" c.close() do:gateway'="" gateway.%Disconnect() quit
}
Run R expression from IRIS SQL
do ##class(Demo.RGateway).FromSQLOnIris()
output :
"mean = 0.5239920995930093"
"median = 0.545"
code :
ClassMethod FromSQLOnIris() { set gateway = ##class(%Net.Remote.Gateway).%New() set tSC = gateway.%Connect("r",55554) if $$$ISERR(tSC) quitset c = ##class(%Net.Remote.Object).%ClassMethod(.gateway,"com.intersystems.rgateway.Helper","createRConnection") // SQL Query set q = "select length from Test.Data" set resultSet = ##class(%SQL.Statement).%ExecDirect( , .q) // Prepare R vector do c.eval("l = c()") while resultSet.%Next() { // Append resultSet to the R vector do c.assign("x", ##class(%Net.Remote.Object).%New(gateway, "org.rosuda.REngine.REXPDouble", resultSet.Length)) do c.eval("l = append(l, c(x))") } // Produce mean from R zw "mean = " _c.eval("mean(l)").asString() // Produce median from R zw "median = "_c.eval("median(l)").asString() do:c'="" c.close() do:gateway'="" gateway.%Disconnect() quit
}
Run R Script and plot
do ##class(Demo.RGateway).FromCSVOnRServe()
output :
"mean = 0.5239920995930093"
"median = 0.545"
"mode = 0.55"
"plot produced in /tmp/data/hist.png"
"plot produced in /tmp/data/hist.jpeg"
"plot produced in /tmp/data/hist.pdf"
code :
ClassMethod FromCSVOnRServe() { set gateway = ##class(%Net.Remote.Gateway).%New() set tSC = gateway.%Connect(..#JGWHOST, ..#JGWPORT) if $$$ISERR(tSC) quitset c = ##class(%Net.Remote.Object).%ClassMethod(.gateway,"com.intersystems.rgateway.Helper","createRConnection", ..#RHOST, ..#RPORT) // Read a file from R // assign an R variable for the data file set filename = "/tmp/data/abalone.csv" do c.assign("filename", filename) // Read the file from the R server do c.eval("data = read.csv(filename)") zw "mean = "_c.eval("mean(data$Length)").asString() zw "median = "_c.eval("median(data$Length)").asString() // Call function from R script // assign an R variable for the script file do c.assign("rFile", "/tmp/r/src/test.R") do c.eval("source(rFile)") zw "mode = "_c.eval("getMode(data$Length)").asString() // Plot graphs // assign a directory to save files do c.assign("dir", "/tmp/data") // produce plot PNG do c.eval("createHistPNG(data$Rings, dir, ""Rings"")") zw "plot produced in /tmp/data/hist.png" // produce plot JPG do c.eval("createHistJPG(data$Rings, dir, ""Rings"")") zw "plot produced in /tmp/data/hist.jpeg" // produce plot PDF do c.eval("createHistPDF(data$Rings, dir, ""Rings"")") zw "plot produced in /tmp/data/hist.pdf" do:c'="" c.close() do:gateway'="" gateway.%Disconnect() quit
}
Another R Gateway
You can have a look at this R Gateway for IRIS :
https://github.com/intersystems-community/RGateway
This one doesn’t use the java gateway and use a direct connection between IRIS and R Server.
Exemple :
Set c = ##class(R.RConnection).%New() // Create a R client
Set x = ##class(R.REXPDouble).%New(3.0) // A single double value
Do c.assign("x", x) // Assign the value to R variable x
Do c.eval("y<-sqrt(x)") // Evaluate R script
Set y = c.get("y") // Get the value of R variable y
Write y.toJSON().%ToJSON()

